EMAN2 Tomography Workflow Tutorial
- This tutorial is best suited for EMAN2 built after 09/27/2018. Not everything described in the tutorial was functioning yet in the 2.22 release.
- The pixel size in the header of the files are incorrect as provided by EMPIAR. The correct Apix value (2.62) should be specified when importing the images.
- To cite:
- Chen, M., Bell, J.M., Shi, X. et al. A complete data processing workflow for cryo-ET and subtomogram averaging. Nat Methods 16, 1161–1168 (2019)
Documentation of some newly developed tools can be found in TomoMore (frequently updated).
There is now a newer pipeline for integrated subtomogram and subtilt refinement. Some documentation can be found in TomoNew (frequently updated).
Contents
-
EMAN2 Tomography Workflow Tutorial
- Computer Requirements
- Download Data
- Prepare input files (~2 minutes)
- Tiltseries Alignment and Tomogram Reconstruction (20 min)
- CTF Estimation (10 min)
- Tomogram evaluation (optional)
- Tomogram annotation (optional)
- Particle picking (10-15 min)
- Particle extraction (a few min)
- Initial model generation (10 - 60 min)
- Template matching (5 min)
- Particle extraction (~1 hour)
- Subtomogram refinement (~6 hr)
- Subtilt refinement (~32 hr)
- Refinement evaluation (optional)
Computer Requirements
- tomographic data processing is normally completed on high-end workstations, not laptops. To complete the tutorial on a laptop you will need to use a significantly reduced data set
- The time estimates for each step are from a workstation with the following configuration:
- Threadripper, 32 core (2990WX)
- 128 GB RAM (64 or perhaps 32 GB would suffice)
- 250 GB free disk space
high performance disk (RAID 5 array or SSD capable of >1 GB/s)
- disk speed has a major impact on performance in many steps
Download Data
This tutorial uses data from EMPIAR: EMPIAR 10064 (the 4 mixed CTEM tilt series)
Prepare input files (~2 minutes)
- Make a new empty folder for the project and 'cd' into that folder
- Make sure any EMAN2 commands you run are executed from within this folder (not any subfolder)
- You may use "Edit Project" from the Project menu to set default values for the project. While not required, it reduces later errors.
- Make sure the workflow mode is set to "TOMO" not "SPR"
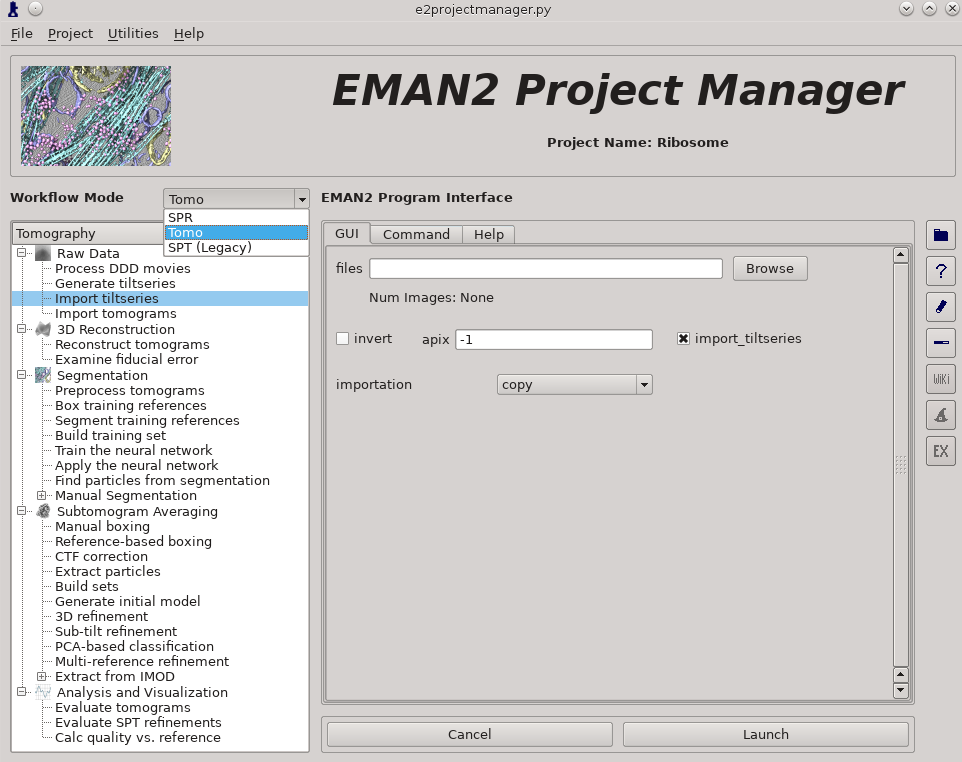
Raw Data -> Import tilt series
Select the files, and make sure importation says copy
In this step you should enter the correct A/pix in the apix box. For EMPIAR10064, this is 2.62. For your own data, you need to know this number.
Once the options are set, press Launch
It is critical that the filenames not contain any spaces (replace with underscore) or periods (other than the final period used for the file extension). "" (double underscore) is also reserved for describing modified versions of the same file, and should not be used in your original files.
For your own data:
If you start from files individual micrographs of the tilt series (after motion correction), use Generate tiltseries to build tilt series from the micrographs. You can build tilt series one by one by selecting all micrographs for one tilt series in tilt_images, specify output and click Launch.
One alternative and easier way is to have all the micrographs in a folder called micrographs, in the same Generage tiltseries panel, put the micrographs folder in tilt_images, check guess and click Launch.
- In principle, the program will guess which files correspond to one tilt series, as well as their tilt angle, from the naming convention of the files. It works most of the time for micrographs produced by major data collection software (SerialEM, EPU, etc.). In the cases it does not work, report to us or use the manual way.
This will create a virtual stack (.lst file) for each tilt series to save disk space. Make sure to always include the micrographs folder in the same directory when moving files around.
Tiltseries Alignment and Tomogram Reconstruction (20 min)
Alignment of the tilt-series is performed iteratively in conjunction with tomogram reconstruction. Tomograms are not normally reconstructed at full resolution, generally limited to 1k x 1k or 2k x 2k, but the tilt-series are aligned at full resolution. For high resolution subtomogram averaging, the raw tilt-series data is used, based on coordinates from particle picking in the downsampled tomograms. On a typical workstation reconstruction takes about 4-5 minutes per tomogram.
For the tutorial tilt-series:
3D Reconstruction -> Reconstruct Tomograms
check alltiltseries
alternatively you can select one or more tilt series from the tiltseries folder
check correctrot
tltstep = 2
clipz = 96
If you wish to look at the intermediate aligned tilt-series and other files, uncheck notmp
- This is not required for the remaining steps in the tutorial, but can be used to help understand how the tomogram alignment works. This requires significant additional disk space. You may consider doing this for only one tomogram.
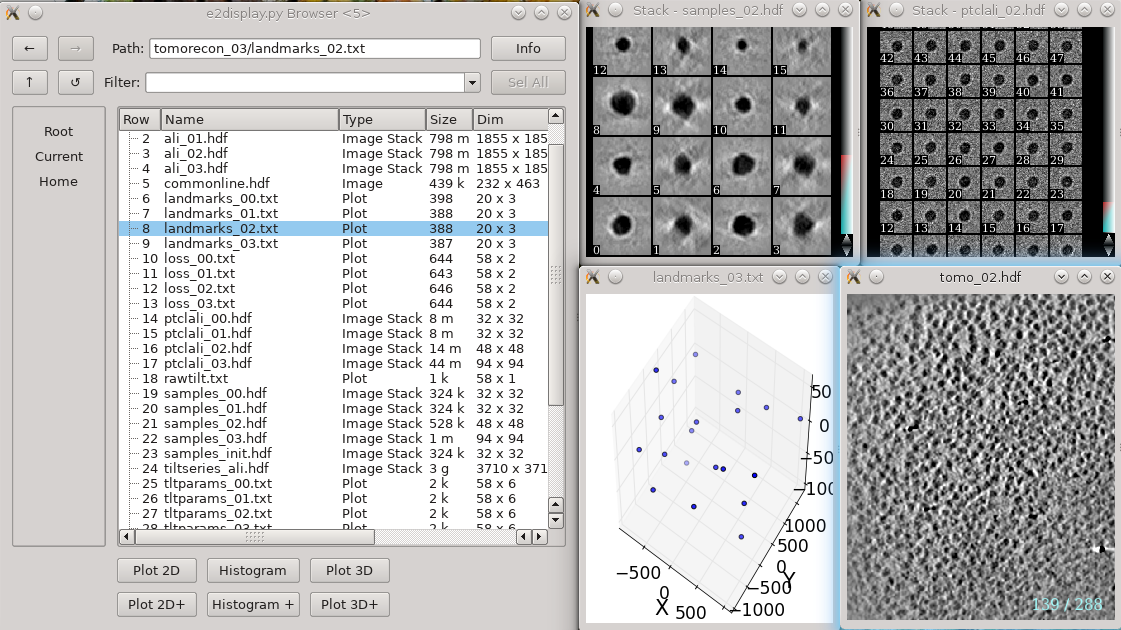
In each tomorecon_XX folder
landmark_0X.txt has the location of the landmarks (which may be fiducials if present) in each iteration
samples_0X.hdf shows the top and side view of those landmarks
ptclali_0X.hdf has the trace of each landmark throughout the tilt series (they should stay at the center of image all the time if the alignment is good)
tomo_0X.hdf is the reconstruction after each iteration
- Launch
For your own data:
Either specify the correct tltstep if the tilt series is in order from one extreme to the other, or specify the name of a rawtlt file (as produced by serialem/IMOD).
While the program can automatically compute the orientation of the tilt axis, it can lead to a handedness ambiguity in the tomogram (it happens to be correct in the tutorial data). For your own data, it is recommended to confirm the handedness in a few good tomograms, then provide the correct tltax value for the reconstruction of all tomograms. To determine the handedness computationally, try the tutorial here for EMAN2 build after 05/23/2019 (or EMAN>=2.31).
In most cases, the default npk should be fine. If fiducials are present, it is not necessary to adjust this number to match the number of fiducials. The program will use any high contrast areas it finds as potential landmarks.
bytile should normally be selected, as it will normally produce better quality reconstructions at higher speed. If 2k or larger tomograms are created, memory consumption may be high, and you should check the program output for the anticipated RAM usage.
- The graphical interface only permits 1k or 2k reconstruction sizes, although 4k reconstruction is supported via the command line. In our experience, 1k/2k is normally sufficient for segmentation or particle picking.
When the sample is thick, some grid-like tiling pattern can be seen in the reconstruction. Checking extrapad can largely reduce the artifacts. In versions after 2/3/2020, there is also a moretile option that further eliminates them. Note these artifacts will NOT impact the subtomogram averaging results because the particles are extracted in a separate process. Checking these options can make the reconstruction process more memory consuming, and up to 5 times slower.
When the sample is thin (purified protein, not cells), it is useful to check correctrot to automatically position tomograms flat in ice
It can also be helpful with thin ice to specify a clipz value to generate thinner tomograms (perhaps 64 or 96 for a 1k tomogram).
xdrift may help a lot when there is significant drift in the tilt series, but it may have worse performance without fiducial.
CTF Estimation (10 min)
For the tutorial tilt-series:
Subtomogram Averaging -> CTF estimation
check alltiltseries
Double check the voltage and cs
- Launch
When working with your own data:
The first two options, dfrange and psrange indicate the defocus and phase shift range to search. They take the format of “start, end, step”, so “2, 5, .1” will search defocus from 2 to 5 um with a step size of 0.1. Units for phase shift is degrees.
For images taken with volta phase plate, we usually have dfrange of “0.2,2,0.1” and psrange of “60,120,2”.
Note that this program is only estimating CTF parameters, taking tilt into account. It is not performing any phase-flipping corrections on whole tomograms. CTF correction is performed later as a per-particle process. This process requires metadata determined during tilt-series alignment, so it cannot be used with tomograms reconstructed using other software packages.
Tomogram evaluation (optional)
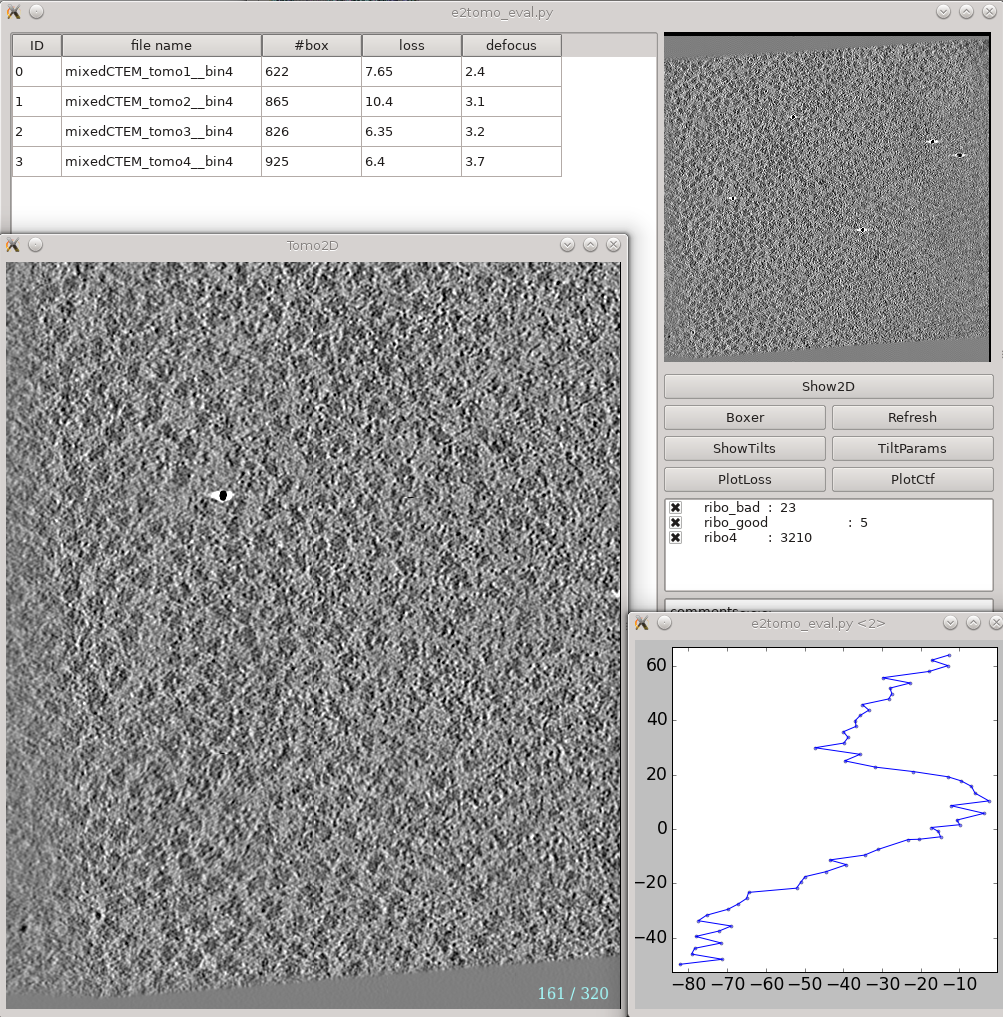
Analysis and visualization -> Evaluate tomograms can be used to evaluate the quality of your tilt series alignments and tomogram reconstructions. This tool will show more information as you progress through the tutorial, but can be used already at this point to make various assessments of your tomograms.
- On the left is a list of tomograms in the project.
- Clicking the header of any column will sort the table by that attribute.
#box is the number of boxes in the tomogram
loss is the average landmark uncertainty in nm. You should not try to compare this number to, for example, the fiducial alignment error in IMOD, as it is computed in a very different way. This number can be useful to detect specific tilt series within a project which have problems, but the absolute number is not a useful value to report/analyze. Even if this number is >5 nm, it is still quite possible to achieve a subnanometer resolution average.
defocus is the average defocus of the tilt series.
- On the right
- The image at the top is the central slice through the tomogram
the show2d button displays the selected tomogram slice-wise.
ShowTilts shows the corresponding raw tilt series
- Please note that most tomograms include some out-of-plane tilt (the actual rotation isn't a simple tilt along a single axis), which is taken into account during alignment. This may make it visually appear that the tilt series alignment is not as robust as it actually is.
Boxer calls the 3D boxer
PlotLoss will plot the fiducial error for each tilt
PlotCtf plot the defocus and phase shift at the center of each tilt image
Tiltparams is a bit more complicated. It plots a point list with 6 columns and a number of rows corresponding to the images in the selected tilt series. These are the alignment parameters for the tilt series.
You can adjust X Col and Y Col in the plot control panel (middle click the plot). The columns represent:
- 0 - tilt ID
- 1 - translation along x
- 2 - translation along y
- 3 - tilt angle around y
- 4 - tilt angle around x
- 5 - tilt angle around z
- The first panel below the buttons are the types of particles and how many of that type are in the project
- The last box is reserved for comments for each tomogram. You can fill in any comments you have on a specific tomogram and it will be saved for future reference.
Tomogram annotation (optional)
In EMAN2 build after 02/01/2020, a new tool is implemented for CNN guided automated particle selectin from tomograms. Check out the guide here.
- Since the tutorial data set is purified ribosomes, this step can be skipped for the tutorial data, and you can move on to template-based particle picking. For cells or other types of complex specimens, tomogram annotation can be used to produce locations of different types of objects.
This section is brief and is only an update to the more detailed tutorial: TomoSeg. Some directory structure and user interfaces have changed in the latest version to match new tomogram workflow as described here:
Segmentation -> Preprocess tomogram
- This step is not always necessary for tomograms reconstructed in EMAN2, but may slightly improve results.
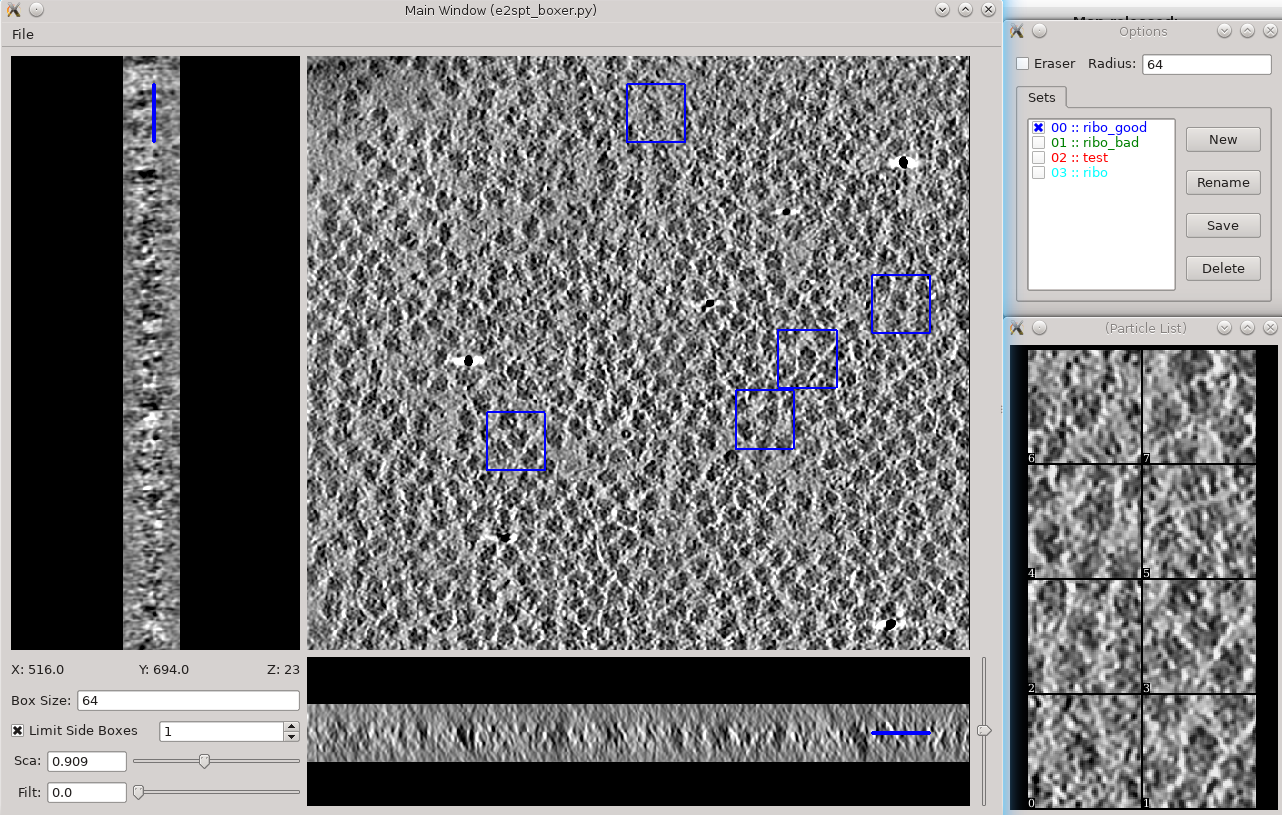
Segmentation -> Box Training References
- This is a newer interface than previously used for this step. Select a few "Good" (regions containing the feature of interest) and "Bad" (regions not containing the feature of interest) boxes.
- "~" and "1" on the keyboard can be used to move along the Z axis.
- The new interface permits different types of features to be identified in a single session and in the same tomogram.
If the different features of interest have very different scale, it is always better to keep the box size at 64, and instead rescale the tomogram. As long as the rescaling is done using EMAN2 utilities, the program will correctly keep track of the geometry relative to the original tomogram & tilt series.
- if you are doing this with the tutorial data, you would only have 2 classes of particles "ribo_good" and "ribo_bad".
When pressing Save all visible particles (box checked next to the class name) will be saved
The rest of the annotation process remain unchanged from the original tutorial, except that now, all trained neural networks and training results are saved in the neuralnets folder, and all segmented maps are in the segmentations folder. You now only specify the label of the output file instead of the full file name.
Segmentation -> Find particles from segmentation to turn segmented maps into particle coordinates.
- Input both the tomogram and its corresponding segmentation, and the particles coordinates will be written into the metadata file.
- Slightly tweaking the threshold parameters may yield better results.
featurename will become the label of particles generated. Those particles can be viewed in the particle picking step and processed in the following protocols.
Particle picking (10-15 min)
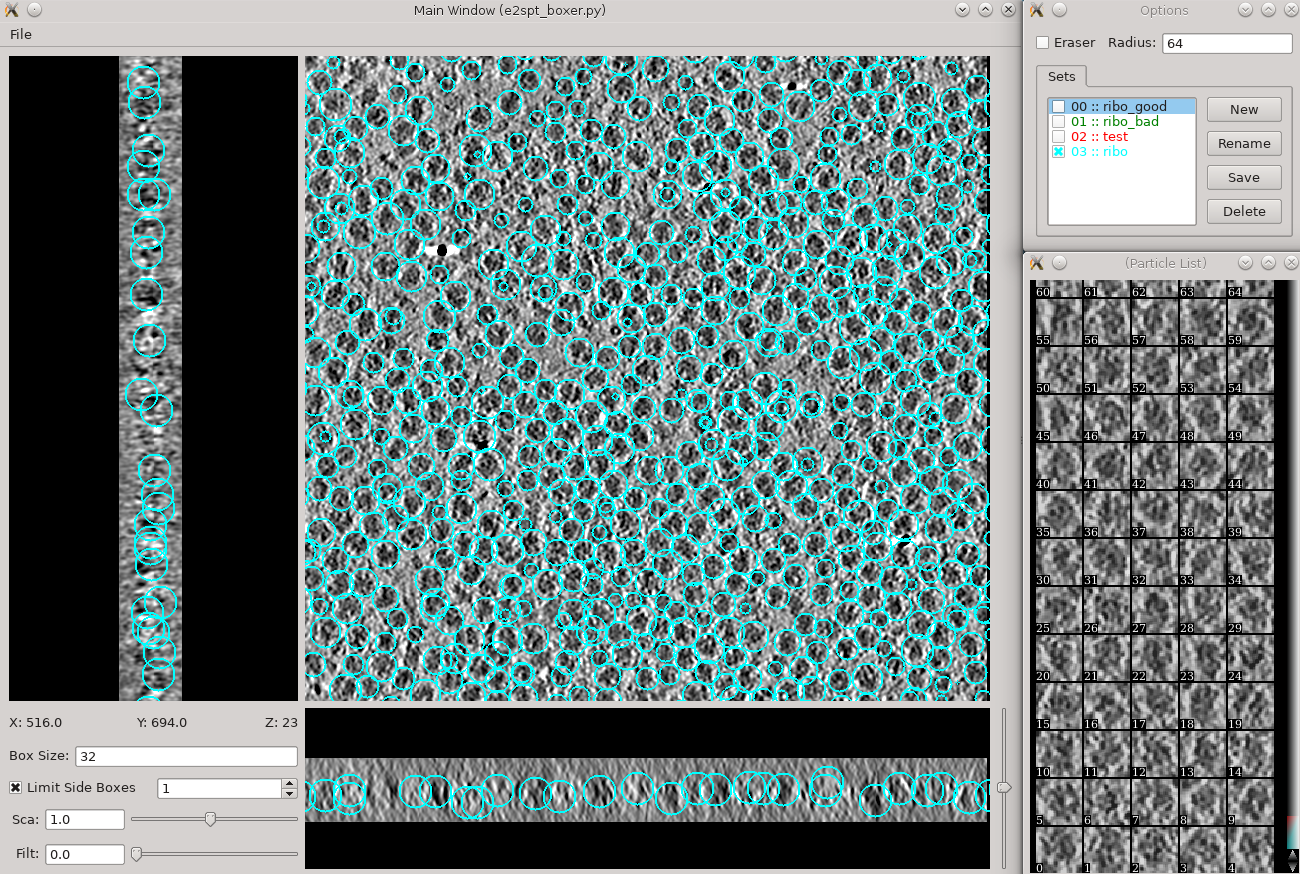
Subtomogram averaging -> Manual boxing Time above is to manually select 30-50 reference particles.
rename the set of boxes to "initribo". This will be used as the label in later stages.
Go through slices along z-axis using ‘~’ and ‘1’ on the keyboard
It will be much easier to locate particles if you adjust the Filt slider to ~70
- left click and drag to place and reposition boxes in any of the 3 views
- Hold down Shift when clicking to delete existing boxes.
- Boxes are shown as circles, which vary in size depending on the Z distance from the center of the particle.
- The interface supports different box types within a single tomogram. Each type has a label. Make sure the label is consistent if selecting the same feature in different tomograms.
- The box size can be set in the main window at the left bottom corner, for the tutorial, use 48 for ribosomes (the unbinned box size is 192).
- If you skipped the tomogram annotation step, we will pick a few particles here to generate an initial model first, and use the initial model as a reference for template matching.
- Select 30-50 particles from a tomogram, then close the boxer window.
- If you have the particle coordinates from tomogram annotation above, you may still wish to do this step to delete any obviously bad particles.
- While you can save 3D particles from the GUI, there is no need to do that here. When you are satisfied with the result, simply close the window.
- You should have ~3000 particles from the 4 tomograms in the dataset.
Particle extraction (a few min)
In this pipeline, the full 1k or 2k tomograms are used only as a reference to identify the location of the objects to be averaged. Now that we have particle locations, the software returns to the original tilt-series, extracts a per-particle tilt-series, and reconstructs each particle in 3-D independently.
For the tutorial tilt-series:
Subtomogram Averaging -> Extract Particles
check alltomograms
set boxsz_unbin to 192.
- If you had the correct size in the previous step this may not be necessary, but it doesn't hurt.
- enter the label you used when picking particles ("initribo" if you followed the instructions above)
- Launch
Subtomogram Averaging -> Build Sets
check allparticles
- Launch
- This will generate particle sets, which are virtual particle stacks that consist of particles with the same label from different tomograms.
For your own data:
If you have gold fiducials present in your tilt series, removing them from the extracted particles/subtilts is critical to success. This can be done using the rmbeadthr option when extracting particles, but a good threshold value must be identified. In cells, a value of 0.5 - 1 is typical, and for isolated particles 1-1.5 may be better. To determine a value rather than just guessing:
extract subtilts for a representative tomogram without using the rmbeadthr option
open one of the subtilts containing one or more fiducials using e2filtertool.py (or pressing the corresponding button in the browser) (see: EMAN2/Programs/e2filtertool)
- configure a Gaussian lowpass filter with cutoff_freq set to 0.01 (100 A) and a Gaussian highpass filter with cutoff_pixels set to 3
By adjusting the min/max values for the image display, you should find a value which shows only the fiducials. That is, adjust min until everything in the images become black except for the fiducials. The min value is the rmbeadthr value to use.
If the box size is correct when you select particles from the GUI, you can leave boxsz_unbin as -1, so the program will keep that box size (scaled to the original tilt series)
If your particles are deeply buried in other densities, using a bigger padtwod may help, but doing so may significantly increase the memory usage and slow down the process.
With CTF information present, it generally does not hurt to check wiener, which filters the 2D particles by SSNR before reconstructing them in 3D.
Specify a binning factor in shrink to produce downsampled particles if your memory/storage/CPU time is limited, but it will also limit the resolution you can achieve.
Initial model generation (10 - 60 min)
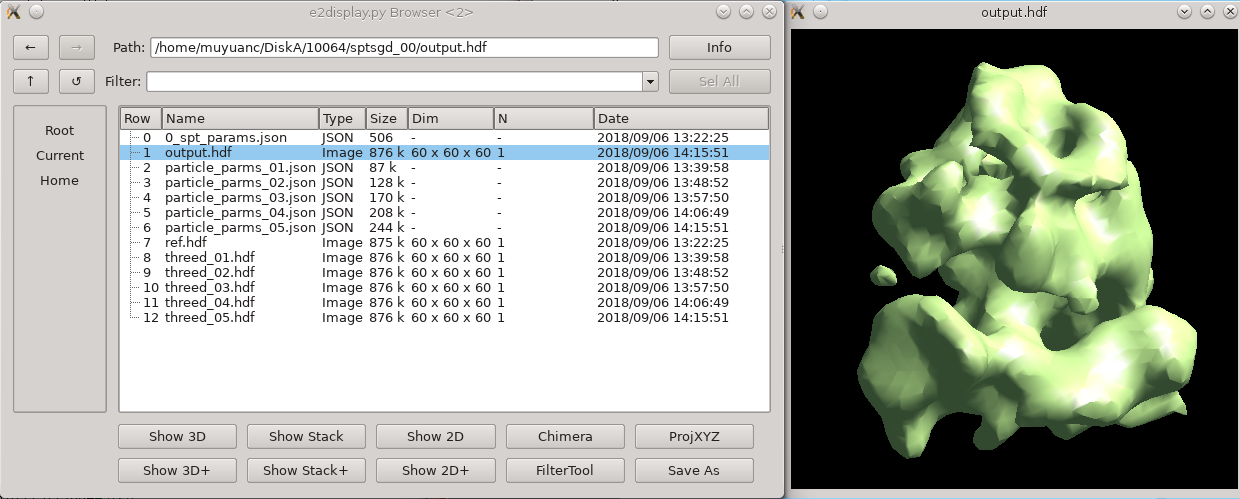
While intuitively it seems like, since the particles are already in 3-D, that the concept of an "initial model" should not be necessary. Unfortunately, due to the missing wedge, and the low resolution of one individual particle (particularly from cells), it is actually critical to make a good starting average, and historically it has been challenging to get a good one, depending on the shape of the molecule. This new procedure based on stochastic gradient descent has proven to be quite robust, but it is difficult for the computer to tell when it has converged sufficiently. For this reason, the default behavior is to run much longer than is normally required, and have a human decide when it's "good enough" and terminate the process. If you use a small shrink value and let it run to completion, it can take some time to run, but this is normally a waste.
For the tutorial tilt-series:
Subtomogram Averaging -> Generate Initial Model
particles should be set to the sets/ribo.lst file you just created (whatever name you used).
set shrink to 2, 3 or 4
- 2 will run slowly but will produce a more detailed initial model (not really necessary)
increasing batchsize will use more cores (if you have more than 12), and may cause it to converge to the correct answer in fewer iterations, but each iteration will not become faster.
The default niter of 5 is typically much more than is required
- Launch
You can terminate the job as soon as sptsgd_00/output.hdf looks reasonable. If you display the progress monitor (4th icon on the right side of the project manager), you can easily kill the job when you're happy. Usually this will take about 10 minutes for the tutorial data.
For your own data:
If your particle has known symmetry, specify that EMAN2/Symmetry
The symmetry you specify will not be imposed on the map unless you also check applysym, but the map will be rotationally aligned so the symmetry axes are in the correct direction, which will make it easier to apply symmetry in later steps. We do not generally recommend checking this box in this step.
setting shrink to something in the range of 2-4 will make the runtime faster but, depending on the shape, could potentially cause problems.
- using more than the minimal 30-50 particles is fine. If you have a very large set of selected particles, go ahead and use them all. This will not slow the process down at all, since it's stochastic.
it is critical that the full sampling box size of the extracted particles divided by shrink be divisible by 2. If not, the program will crash.
Template matching (5 min)
In this step, we will use the initial model you just produced as a template for finding all of the ribosomes in all 4 tomograms. If you completed the Tomogram Annotation step above, and have already extracted a full set of 1000+ particles, then you can skip this step, as we already have all of the particles. Note that here, and everywhere else in the tomography pipeline, reconstructed particles have positive contrast (look white in projection) and tomograms/tilt series have negative contrast (look dark in projection). If you wish to use a reference volume from the PDB or somesuch, then it should have positive contrast as is normal in the single particle CryoEM field.
Subtomogram Averaging -> Reference Based Boxing
browse to select tomograms. Select all 4 tomograms.
set reference to the output.hdf file you produced in the previous step.
set label to "ribo"
set nptcl to 1000 (the maximum number of particles per tomogram)
IMPORTANT NOTE: with these parameters it is possible to reproduce a subnanometer resolution ribosome structure, but the final refinement could take more than 24 hours to run. If you set nptcl to, say 100 instead of 1000, your resolution will be lower, but the subsequent jobs will complete ~10x faster.
- Launch
when this finishes, you can use the same Manual Boxing tool you used before to look at the particles which were selected. You may wish to manually remove any bad particles it selected. For the tutorial data set or other tomograms of purified protein, this process should work pretty well. For cells you might wish to use the Tomogram Annotation method above.
- note that this process stores 3-D particle locations in the appropriate info/* files, but does not extract particles from the micrographs
Particle extraction (~1 hour)
Again, if you already did Tomogram Annotation above, this step isn't necessary. It is only required if you just did Template Matching.
Since this involves several thousand particles instead of 30-50, it will take quite a lot longer to run. The actual time will depend partially on the speed of your storage.
For the tutorial tilt-series:
Subtomogram Averaging -> Extract Particles
check alltomograms
set boxsz_unbin to 192.
set label to "ribo"
- Launch
Subtomogram Averaging -> Build Sets
check allparticles
- Launch
- This will generate particle sets, which are virtual particle stacks that consist of particles with the same label from different tomograms.
Subtomogram refinement (~6 hr)
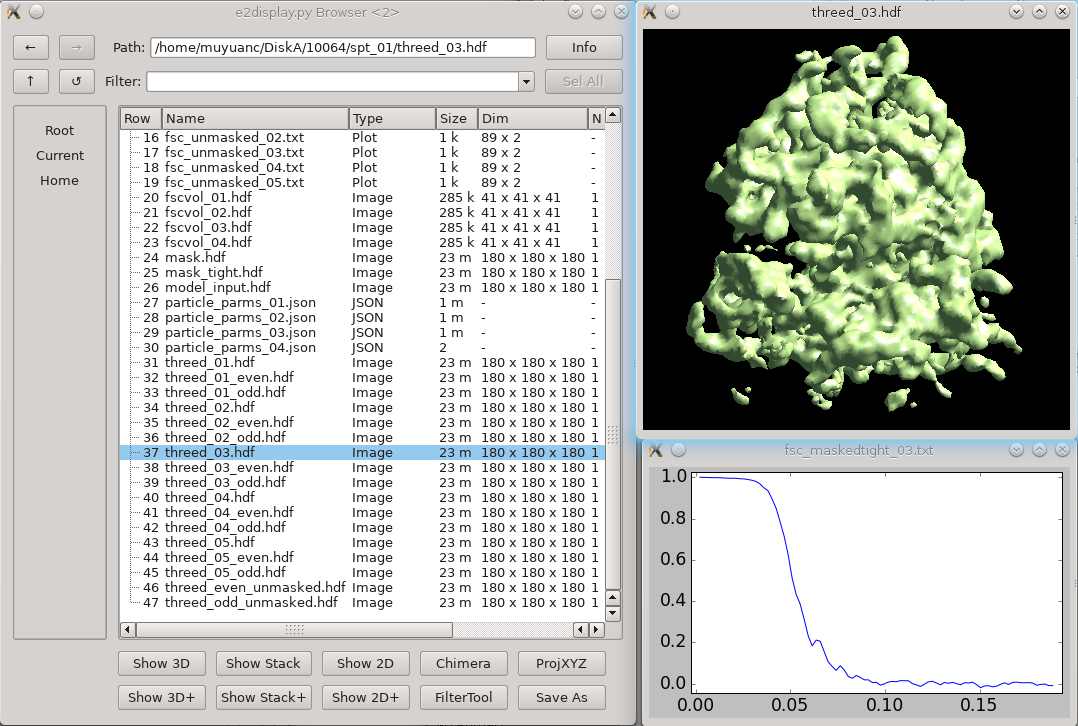
This step performs a conventional iterative subtomogram averaging using the full set of particles. Typically it will achieve resolutions in the 15-25 A range with a reasonable number of particles. As it involves 3-D alignment of the full set of particles multiple times, it takes a significant amount of compute time. Higher resolutions are achieved in the next stage after this (subtilt refinement).
For the tutorial tilt-series:
Subtomogram Averaging -> 3D Refinement
set particles to "sets/ribo.lst"
set reference to "output.hdf" from Initial Model Generation
set goldstandard to 30
set mass to 3000
set threads to the number of CPUs on your machine
- Launch
Results will gradually appear in spt_XX/
For your own data:
- If your molecule has symmetry, you should specify it, but it's important that the alignment reference you provide has been properly aligned to the symmetry axes of whichever symmetry you specify.
localfilter will use e2fsc.py to compute a local resolution map after each iteration and filter the map accordingly. This is useful for molecules with significant variability.
If you suspect that a large fraction of your particles are "bad" in some way, you may wish to try reducing pkeep, which will hopefully exclude bad particles preferentially over "good" particles.
Subtilt refinement (~32 hr)
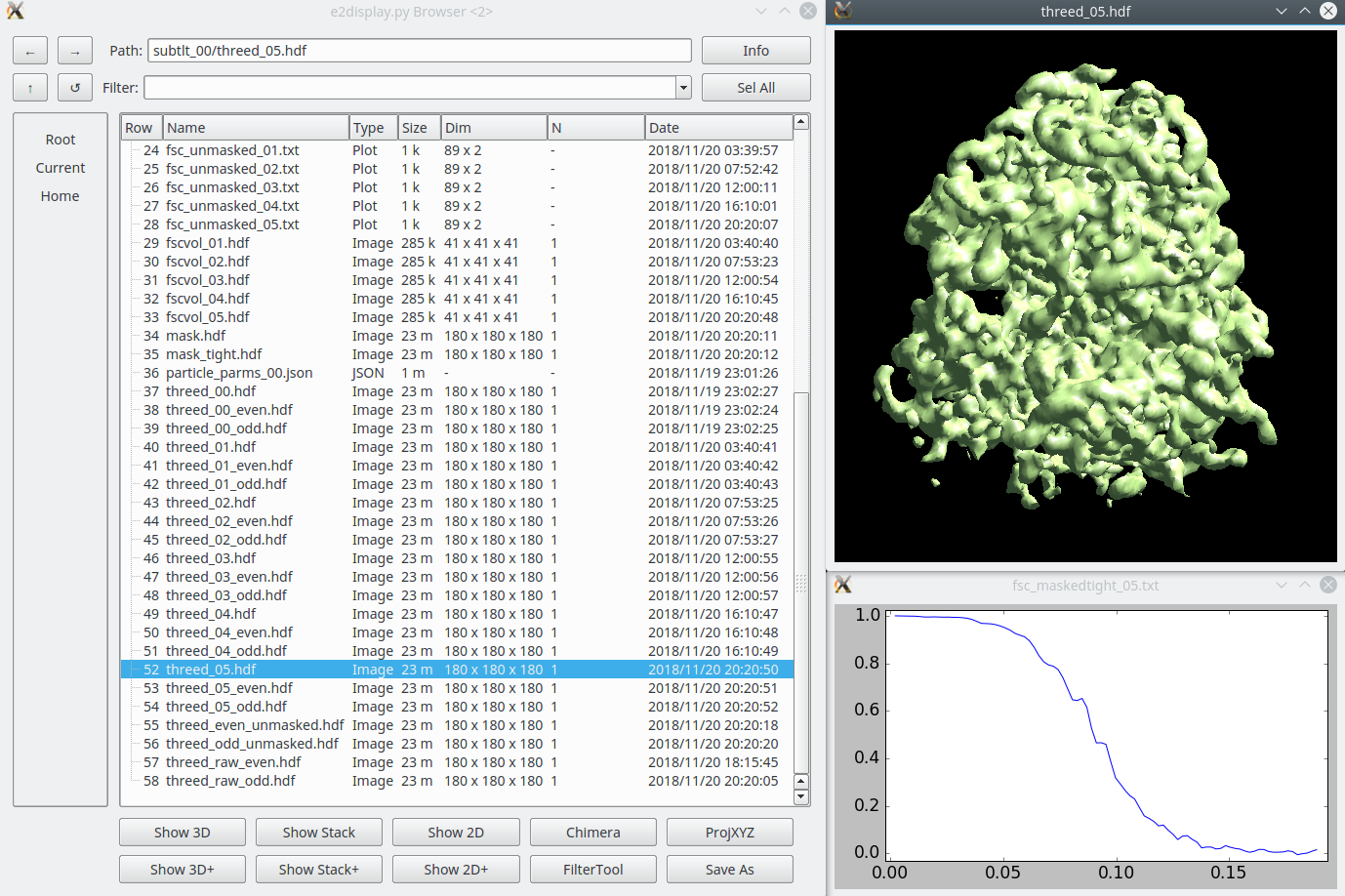
With the results of a good subtomogram alignment/average, we are now ready to switch to alignment of the individual particle images in each tilt, along with per-particle-per-tilt CTF correction and other refinements. This is effectively a hybrid of single particle analysis and subtomogram averaging, and can readily achieve subnanometer resolution IF the data is of sufficient quality. The tutorial data set is, but many cellular tomograms, for example, are not collected with high resolution in mind, and even with this sort of refinement will be unable to achieve resolutions better than 10-30 A, depending on the data. This process is completely automatic, based on all of the metadata collected up to this point. While it is possible to perform "subtomogram refinement" with subtomograms from any tomogram, Subtilt Refinement cannot operate properly unless all preceding steps occurred within EMAN2.
For the tutorial tilt series:
Subtomogram Averaging -> Sub-tilt Refinement
path should be set to the name of one of a "spt_XX" folder to use as a starting point for the refinement
iter can be -1 to use the last complete iteration in the "spt_XX" folder. Alternatively you can specify a specific iteration to use
parallel should be "thread:N" where N is the number of cores you wish to use on a single machine. This job can be run on a linux cluster if you like: EMAN2/Parallel.
threads should also be set to the number of cores to use on a single machine
- Launch
For your own data:
niters is the number of iterations to run. The default of 4 should achieve convergence in most cases.
keep is the fraction of tilt images to use in the final map. This defaults to 0.5, meaning the worst 1/2 of the tilts for each particle will be discarded. This permits tilts which contain, for example, projections of fiducials or other strong densities, or with large amounts of motion to be automatically excluded in the final reconstruction.
maxalt specifies the maximum tilt angle to include from each particle. Most tilt series are collected such that the small tilt angles will have the least radiation damage, and very often high tilts suffer from more motion artifacts. If you enter, for example, "45" in this box then tilts <-45 and >45 will be discarded automatically. In most cases keep will already serve a similar purpose.
Congratulations! The final result of the tutorial will be found in "subtlt_00/". The final 3-D map will be "threed_04.hdf" with the default parameters. The final gold standard resolution curve will be "fsc_maskedtight_04.txt". The optional steps below are tools you can use to evaluate your results in more detail.
Refinement evaluation (optional)
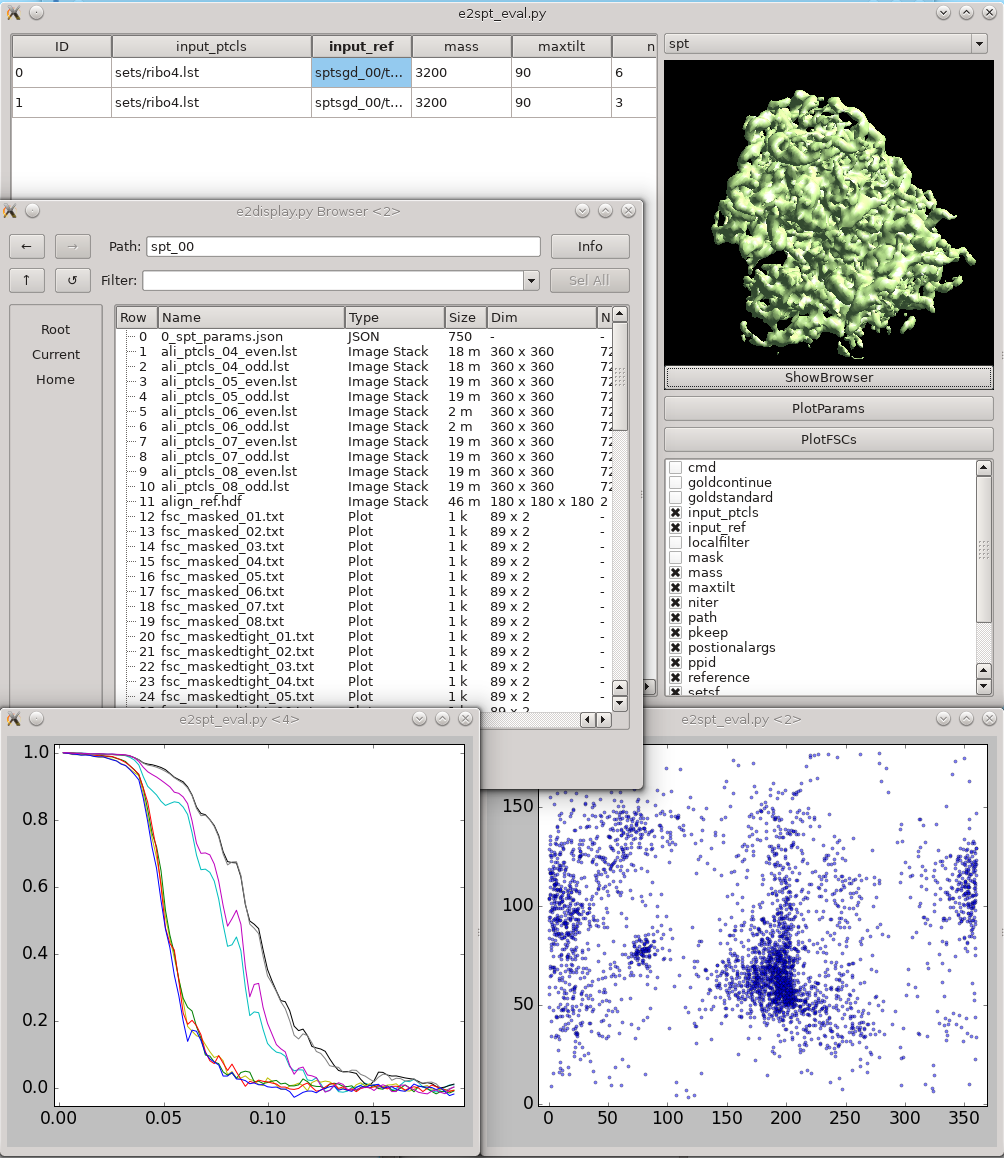 This tool helps visualize and compare results from multiple subtomogram refinement runs.
This tool helps visualize and compare results from multiple subtomogram refinement runs.
Analysis and Visualization -> Evaluate SPT Refinements
In the GUI, you can look at all spt_XX or sptsgd_XX folders and compare the parameters which were used for each, as well as the resulting maps.
- Switch between folder types using the menu at top right.
- Columns can be sorted by clicking on the corresponding header.
- Uncheck items in the list at bottom-right to hide corresponding columns
ShowBrowser will bring up the e2display.py browser in the folder of the selected row.
!PlotFSC will display the "tight" FSC curve over all iterations.
PlotParams will plot the Euler angle distribution and other alignment parameters
- The 8 columns in the plot are:
- 0 - az (EMAN convention Euler angle)
- 1 - alt
- 2 - phi
- 3 - translation in X
- 4 - Y
- 5 - Z
- 6 - alignment score
- 7 - missing wedge coverage
- The 8 columns in the plot are:
