|
Size: 261
Comment:
|
Size: 1426
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
| == e2helixboxer.py: Overview == e2helixboxer.py is used to select rectangular 2D projections of helices from a micrograph, and extract overlapping particles from the boxed helices. |
= e2helixboxer.py: Overview = e2helixboxer.py is used to select rectangular 2D projections of helices from a micrograph, and extract overlapping particles from the boxed helices. The boxing must be done in GUI mode, but extracting particles from boxed regions may be done from the GUI or from the command line. == GUI mode == To start the program's graphic user interface, use the "--gui" option. You can follow this with a micrograph filename, a list of micrograph filenames, or nothing. {{{ $ e2helixboxer.py --gui <micrograph1> <micrograph2> <...> }}} {{{ $ e2helixboxer.py --gui $ e2helixboxer.py --gui 101.mrc $ e2helixboxer.py --gui *.mrc micrograph.hdf *.img abc.dm3 }}} The left window is the helix viewer, the middle is the micrograph viewer, and the right is the main window. |
| Line 6: | Line 19: |
| == GUI mode == | The main window shows a list of open micrographs and how many helices you have boxed in each one. (Actually, only one micrograph is loaded in memory at a time, but this allows for quick switching between particles.) Any micrographs you specified after the "--gui" option will be listed. ||||<tablestyle="width: 485px; height: 212px;">Box Editing|| ||Draw box||Left click and drag|| ||Move box||Left click near box center and drag|| ||Move one box endpoint||Left click near box end and drag|| ||Delete box||Hold shift and left click inside box|| |
e2helixboxer.py: Overview
e2helixboxer.py is used to select rectangular 2D projections of helices from a micrograph, and extract overlapping particles from the boxed helices. The boxing must be done in GUI mode, but extracting particles from boxed regions may be done from the GUI or from the command line.
GUI mode
To start the program's graphic user interface, use the "--gui" option. You can follow this with a micrograph filename, a list of micrograph filenames, or nothing.
$ e2helixboxer.py --gui <micrograph1> <micrograph2> <...>
$ e2helixboxer.py --gui $ e2helixboxer.py --gui 101.mrc $ e2helixboxer.py --gui *.mrc micrograph.hdf *.img abc.dm3
The left window is the helix viewer, the middle is the micrograph viewer, and the right is the main window.
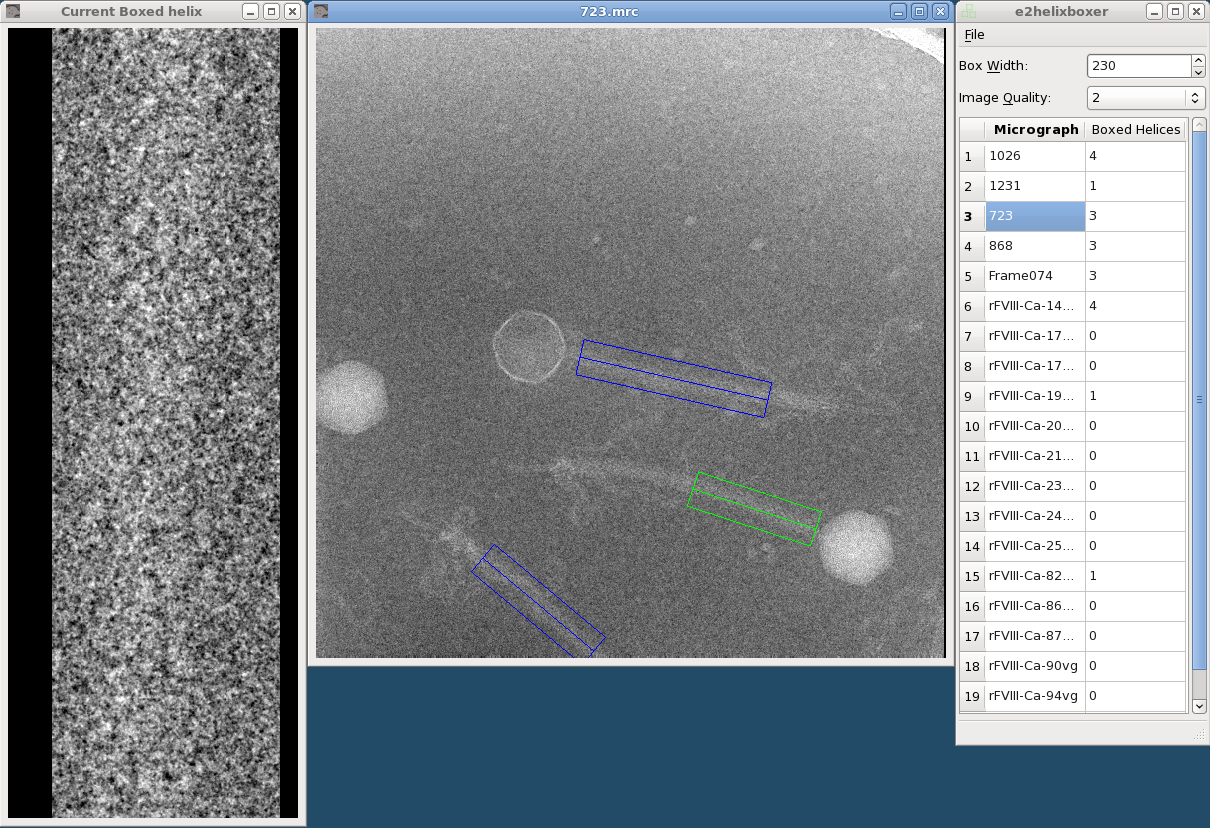
The main window shows a list of open micrographs and how many helices you have boxed in each one. (Actually, only one micrograph is loaded in memory at a time, but this allows for quick switching between particles.) Any micrographs you specified after the "--gui" option will be listed.
Box Editing |
|
Draw box |
Left click and drag |
Move box |
Left click near box center and drag |
Move one box endpoint |
Left click near box end and drag |
Delete box |
Hold shift and left click inside box |
